Do the same thing for the accession number NM_006516 do download GLUT1 RNA 10. Choose right destination in your folder, keep accession number with your naming,like: NG-0088232 Human GLUT3 9. Close the sequence, and the program will ask: Save changes to 'NG_008232' before closing?, choose "Yes 8. DoubleClick on it, the sequence of 40, 802 be will appear in the bottom. This sequence will appear: NG 008232 Homo sapiens solute carrier family 2 member 1 (SLC2A1), RefseqGene on chromosome 1 40802 6. accession number for the human SLC2A1 DNA (NG_008232). In NCBl search site click nucleotide on the top, on the left choose "all field" and copy the Glucose Transporter 1" (SLC2A1). Go Download and click on Search for sequences on NCBI. 
In the left navigation area click on CLD data, highlight it 2. The nonfunctional/defective pre-C protein of this isolate highlights the need for further investigation of this type of HBV in Bangladesh, along with its clinical outcome.CLC SEQUENCE VIEWER 7.7.1 Download this program from the site: cic.sequence-viewer/#download 1. The following nucleotide mutations were found: A2962G and C3026A in the pre-S1 region T1753C, G1757A, and A1762C in the basic core promoter region and G1896A and A2189T in the pre-C/C regions. A vaccine-escape HBsAg A128V mutation was found. The viral isolate was characterized as genotype D (subgenotype D2) and subtype (serotype) ayw3 and was evolutionarily related by phylogenetic analysis to the HBV isolated from Bangladesh, Italy, Indonesia, and Estonia ( Fig. 1). This isolate showed a nonfunctional/defective pre-C protein due to a mutation at nucleotide position G1896A (insertion of stop codon TAG). The G+C content of the genome was 48.65%. The minimum quality value (QV) scores of the reads used for the assembly were 20. The entire assembled genome was circular and 3,182 bp long, with four overlapping ORFs, namely, P, S, C, and X. The newly obtained HBV (strain HBV_BAU1) genome sequence was aligned with different genotypes and subgenotypes for the construction of phylogenetic trees using CLC Sequence Viewer ( 8). 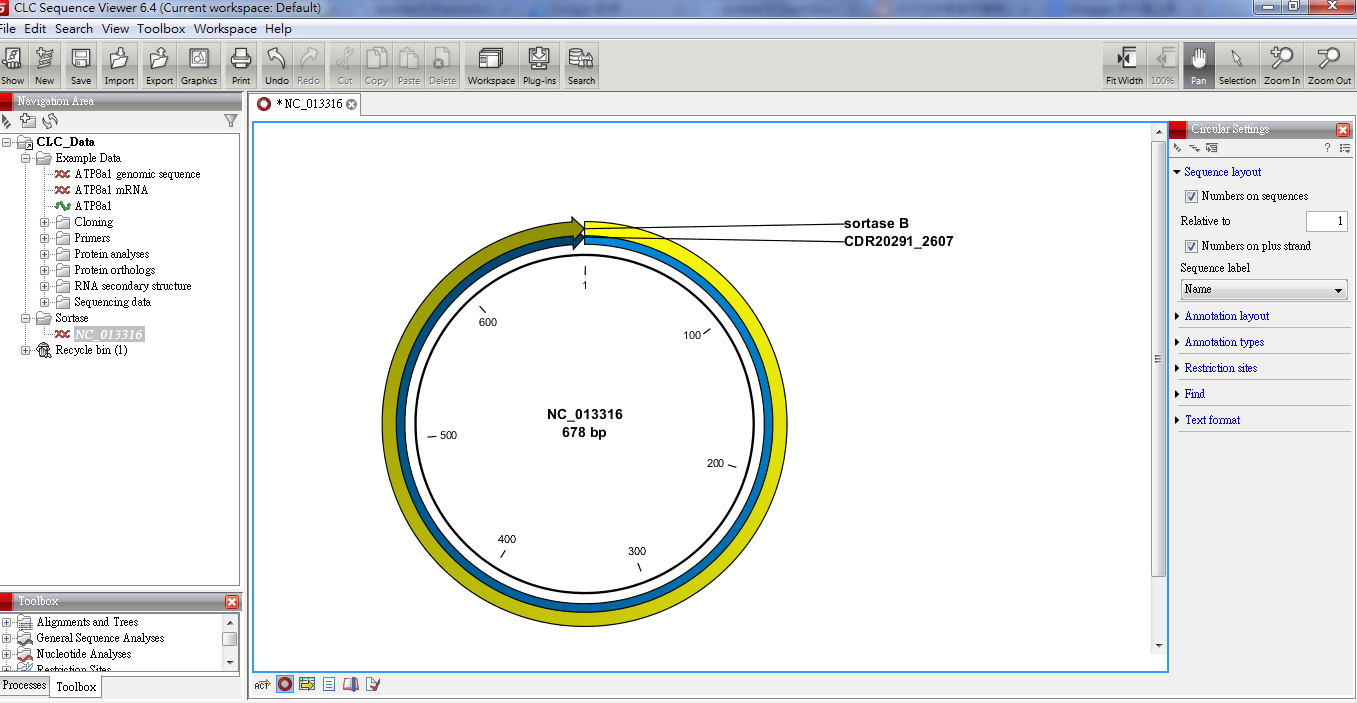
Determination of the genotype and subgenotype as well as the mutation analysis were performed using Geno2pheno:hbv ( ) and HIV-Grade:HBV-Tool ( ) with default parameters. Raw sequence data were edited, annotated, and analyzed using CLC Sequence Viewer. The sequencing reactions were performed according to our previously reported protocol using the BigDye Terminator v3.1 cycle kit and the BigDye Terminator v1.1 and v3.1 5× sequencing buffer (Applied Biosystems) and were sequenced using an Applied Biosystems 3730 DNA analyzer (California, CA, USA) ( 3). 
The PCR products were purified using a MonoFas DNA purification kit (GL Sciences, Inc., Tokyo, Japan) and were used as the templates for sequencing. Amplification of the HBV complete overlapping ORFs P, pre-C, and X and the partial S ORF was carried out using specific primers ( Table 1), as previously described ( 3). Viral DNA from serum samples was extracted using a QIAamp DNA minikit according to the manufacturer’s protocol. HBV was detected in a patient serum sample ( 3). We report the full-genome sequence of a pre-C-defective HBV mutant isolated in Bangladesh from a clinically infected patient and report a mutation analysis. HBV is highly prevalent in Bangladesh, with the genotypes A, C, and D being responsible for 66% of HCC in this country ( 7). Pre-C-defective HBV mutants are infectious but are less sensitive to interferon treatment ( 6). The HBV genome encodes four major open reading frames (ORFs), namely, polymerase, HBsAg, core, and X precore (pre-C)-defective mutations occur due to the introduction of stop codons in this region ( 6). However, the prevalences of HBV genotypes and subtypes vary depending on the geographical region, and genomic variations have been associated with HBV pathogenesis ( 2, 5). Nucleotide sequence variation in the genome has been used to classify HBV into different genotypes (A to J) ( 3, 4). Hepatitis B virus (HBV), a causative agent for acute/chronic hepatitis and hepatocellular carcinoma (HCC), has a circular partially double-stranded DNA genome, is about 3.2 kb long, and belongs to the Hepadnaviridae family and Orthohepadnavirus genus ( 1, 2). 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
